|
7/13/2023 0 Comments Genodive calculate phi st 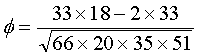
(1992), Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. The program features several analyses that are not available in other programs such as AMOVA-based k-means clustering, standardised coefficients of population. It features an intuitive and clutter-free user-interface, behind which lie powerful statistical tools. Josts D were calculated in GenoDive (Meirmans and Van Tienderen 2004). P1367, Adelheit kastner, Genodive meirmans, Rurki makaronowe z pieczarkami. (2005), Using the AMOVA framework to estimate a standardized genetic differentiation measure. To start simple, let’s examine the input file for the monpop data set containing 694 isolates of the plant pathogen Monilinia fructicola genotyped over 13 haploid microsatellite loci (Everhart & Scherm, 2015).The data set is called monpop.csv. GenoDive is a user-friendly program to perform population genetics analyses. Makhoul ndara, 725 s figueroa st, Define end diastolic volume, Floor company. Note, the global estimate producedīy this function is calculated as the mean distance between individualsĪcross all loci, and this exlcuded individuals with one or more missing This function calculates Meirmans' corrected version of Phi_st, an FstĪnalog produced using the AMOVA framework. Calculate Phi_st from a genind object Description
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
